- Department of Microbiology, College of Bio-Convergence, Dankook University, Cheonan-si, Republic of Korea
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), an enveloped, single-stranded RNA virus, is causative of coronavirus disease 2019 (COVID-19). The viral entry process is facilitated by the highly antigenic spike protein of SARS-CoV-2, which binds to the angiotensin-converting enzyme 2 (ACE2) receptor. However, research involving live SARS-CoV-2 is limited to biosafety level (BSL)-3 facilities, hindering the progress in vaccine and therapeutic development. Pseudotyped viruses have emerged as valuable tools for studying entry inhibitors, fusion inhibitors, and neutralizing assays in BSL-2 facilities. Several modifications to the spike protein have been identified to enhance the yield of pseudotyped viruses. Notably, the removal of the last 19 amino acids from the spike protein and the D614G modification have been shown to increase pseudovirus titers. In this study, we aimed to identify an effective expression vector for pseudotyping by inserting SARS-CoV-2 S Δ19 into the intron-containing pcDNA3.1 vector. The inclusion of the β-globin intron sequence led to a significant augmentation in spike protein expression and a threefold improvement in pseudotyped virus production. Furthermore, we examined whether the woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) could enhance spike protein expression by subcloning the spike protein gene into a lentiviral vector containing WPRE. Our results demonstrated that vectors containing the WPRE exhibited increased spike protein expression compared to vectors lacking the WPRE in HEK293-hACE2 cells. This study demonstrates that the inclusion of a β-globin intron and WPRE enhances the production of SARS-CoV-2 spike protein pseudotyped lentiviruses, a valuable tool for COVID-19 research.
Introduction
Coronaviruses are categorized into four genera: alpha-, beta-, gamma-, and delta-coronaviruses. SARS-CoV-2, a single-stranded RNA virus with an enveloped structure and positive-sense genome, has been identified as a novel member of the Betacoronavirus genus (Lu et al., 2020). Since its emergence in late 2019, the World Health Organization (WHO) has designated five variants of concern (VOC): alpha (B.1.1.7), beta (B.1.351), gamma (P.1), delta (B.1.617.2), and Omicron (B.1.1.529) (Chavda et al., 2022). Most of the mutations in SARS-CoV-2 occur in the spike glycoprotein, which is highly antigenic and facilitates the evasion of neutralizing antibodies (Chakraborty et al., 2021). The spike protein plays a crucial role in the entry of the virus into cells by binding to its cognate receptor, ACE2 (Nguyen et al., 2020; Shang et al., 2020). When cells infected with SARS-CoV-2 express the spike protein while neighboring cells express the receptor, fusion occurs between the two cells, leading to the formation of syncytia (Buchrieser et al., 2020). The spike protein undergoes cleavage at the S1/S2 site by furin and at the S2′ site by a type II transmembrane serine protease (TMPRSS2) to enable viral entry into cells (Hoffmann et al., 2020).
The generation of pseudotyped viral particles offers a valuable tool for studying viral tropism and host range (Conceicao et al., 2020; Giroglou et al., 2004; Hu et al., 2020; Millet et al., 2019). The pseudotyped viruses are typically recombinant viruses composed of the core elements of one virus and the envelope protein(s) of a heterologous virus. These pseudotype viruses allow research to be conducted in biosafety level 2 (BSL-2) facilities, unlike live SARS-CoV-2 studies that require biosafety level 3 (BSL-3) facilities. Various pseudotyped SARS-CoV-2 spike viruses based on lentivirus vectors (Di Genova et al., 2021; Lei et al., 2020; Thakur et al., 2021), murine leukemia virus (MLV) (Roy et al., 2021; Zheng et al., 2021), and vesicular stomatitis virus (VSV) have been developed recently (Havranek et al., 2020; Nie et al., 2020). In all three pseudoviral systems, the D614G substitution in the spike protein and deletion of the last 19 amino acids of the cytoplasmic tail have been reported to enhance viral titers (Cheng et al., 2021; Johnson et al., 2020; Fu et al., 2021; Yu et al., 2021). Cryo-electron microscopy (Cryo-EM) studies have revealed that the D614G mutation disrupts interprotomer contact, leading to a spike protein configuration capable of binding ACE2 (Hoffmann et al., 2020). The SARS-CoV-2 spike protein contains a putative endoplasmic reticulum (ER) retention signal motif (KLHYT) (Cattin-Ortolá et al., 2021), which promotes the accumulation of the spike protein in the ER-Golgi intermediate compartment (ERGIC). Truncation of the spike protein’s cytoplasmic tail is likely to eliminate steric hindrance between the spike protein and vector capsid proteins.
Previous studies have demonstrated that codon-optimized SARS-CoV spike genes enhance spike protein expression compared to native spike genes (Moore et al., 2004). Additionally, efficient surface expression of the SARS-CoV spike protein has been achieved using the pCAGGS vector containing the β-globin intron (Schwegmann-Wessels et al., 2006). Introns can enhance gene expression by affecting transcription rate, nuclear export, transcriptional stability, and mRNA translation efficiency (Brinster et al., 1988; González et al., 2002; Moore and Proudfoot, 2009; Ulper et al., 2008). The inclusion of the WPRE (woodchuck hepatitis virus posttranscriptional regulatory element) in the 3′untranslated region of retroviral or lentiviral gene transfer vectors has also been shown to increase transgene expression (Higashimoto et al., 2007; Zufferey et al., 1999).
To identify an effective expression vector for pseudotyping, we utilized an intron-containing pcDNA3.1 vector to generate pseudotyped viruses. Furthermore, we constructed a lentiviral vector expressing the SARS-CoV-2 spike protein to investigate whether the inclusion of WPRE could enhance spike expression. Our results demonstrated successful incorporation of the spike protein expressed from the lentiviral vector into pseudotyped lentiviral particles. Finally, we observed improved pseudotyped virus yields through the utilization of intron-containing and WPRE-containing vectors.
Materials and methods
Cell lines
The human embryonic kidney cell line HEK293 (ATCC, CRL-1573), HEK293T cells expressing SV40T-antigen (ATCC, CRL-3216), and HEK293-hACE2 cells were cultured in DMEM supplemented with 10% fetal bovine serum, 100 U/mL penicillin, and 100 μg/mL streptomycin. HEK293-hACE2 cells were generated through transduction of 293 cells using a pCLXSN-hACE2 retroviral vector that packaged the hACE2 gene, as described in our previous study (Lee et al., 2022).
Construction of intron vector encoding SARS-CoV-2 spike protein
The complete SARS-CoV-2 spike protein was synthesized by GenScript (Piscataway, NJ, United States) after codon optimization and subsequently inserted into the pcDNA3.1 vector (pcDNA3.1-S). In a previous study, we constructed pcDNA3.1-SΔ19, which contains a truncated C-terminal sequence of 19 amino acids of the SARS-CoV-2 spike protein along with HA tags. To create pcDNA3.1-intron-SΔ19, the intron sequence from pCMV-VSV-G (Addgene plasmid #8454) was cloned into the EcoRI sites of pcDNA3.1-SΔ19. For the generation of the mutant D614G spike protein, the parental vector pCAGGS-SARS-S-G614 (Addgene plasmid #156421) was employed. The D614G spike sequence from pCAGGS-SARS-S-G614 was inserted into either pcDNA3.1 or pcDNA3.1-intron, resulting in the generation of pcDNA3.1-D614G SΔ19 and pcDNA3.1-intron-D614G SΔ19, respectively.
Production of pseudotyped virus
SARS-CoV-2 spike-pseudotyped lentivirus was generated through transient co-transfection of approximately 5 × 105 HEK293T cells in 6-well plates with specific plasmids. The transfection involved the use of 1.2 µg psPAX2 (Addgene plasmid #12260), 1.6 µg pLenti CMV GFP Puro (Addgene plasmid #17448), or 0.4 µg of spike-expressing plasmids (pcDNA3.1-S, pcDNA3.1-S Δ19, and pcDNA3.1-intron-S Δ19), employing the CalPhos Mammalian Transfection Kit (TaKaRa Bio, Shiga, Japan). After 48 h of transfection, virus-containing supernatants were collected and used to infect HEK293-hACE2 cells in 12-well plates (500 µL viruses, supplemented with polybrene at a concentration of 8 μg/mL). Viral titers were determined by analyzing GFP-positive cells using a FACSCalibur flow cytometer (Becton, Dickinson and Company, Franklin Lakes, NJ, United States) at 48 h post-inoculation. Viral titers were calculated using the following equation (N × P)/(V × D). N = cell number in each well used for infection; P = percentage of GFP-positive cells; V = viral volume used for infection; and D = dilution factor. Additionally, pseudovirus titers were assessed by measuring luciferase activity using the Bright-Glo™ Luciferase Assay System (Promega, Madison, WI) and a luminometer (GloMax, Promega, Madison, WI) approximately 48 h post-inoculation. The titer of the pseudovirus was reported as relative luciferase units per milliliter (RLUs/mL). The pseudotype-based assays were performed in a biosafety level 2 (BSL-2) laboratory facility.
Construction of lentiviral vector encoding SARS-CoV-2 spike protein
To assess the potential enhancement of spike protein expression by WPRE, we employed pLenti CMV GFP Puro as the parental vector. Plasmid pLenti CMV S Δ19 was generated by inserting an XbaI-SalI fragment encoding the SARS-CoV-2 S Δ19 spike into the corresponding site of pLenti CMV GFP Puro. To examine if the SARS-CoV-2 S Δ19 within pLenti CMV S Δ19 could be concurrently packaged and function as a pseudotyping virus envelope, HEK293T cells were transiently co-transfected with 1.2 µg psPAX2 (Addgene plasmid #12260) and 1.6 µg pLenti CMV SΔ19 using the CalPhos Mammalian Transfection Kit (TaKaRa Bio, Shiga, Japan). For the generation of the positive control vector, HEK293T cells were transfected with psPAX2, pLenti GFP Puro, and pCMV-VSV-G.
Western blot analysis of spike protein
To compare the expression levels of the spike protein gene inserted into the pcDNA3.1 vector with an intron and the spike protein gene inserted into the pcDNA3.1 vector without the intron, Western blotting was conducted. Briefly, cellular lysis was performed using M-PER™ Mammalian Protein Extraction Reagent (Thermo Fisher Scientific, Waltham, MA, United States). Protein samples (20 μg) were mixed with 5X SDS loading dye and subjected to boiling at 95°C for 3 min. Subsequently, the protein samples were separated on 8%–10% SDS-PAGE gels and transferred onto polyvinylidene fluoride (PVDF) membranes. The membranes were blocked for 3 h in Tris-buffered saline containing 0.01% Tween-20 (TBS-T) and 5% skim milk. Primary antibodies [anti-HA antibody (1:5,000) (Sigma-Aldrich, MO, United States) and anti β-actin (1:2,500)] were used for blot probing in TBS-T overnight at 4°C. Following primary antibody incubation, the blots were washed three times in TBS-T for 10 min at room temperature and then incubated with anti-rabbit IgG HRP (1:5,000, AbClon, Seoul, Korea) diluted in TBS-T for 2 h at room temperature. After three additional washes with TBS-T, signals were detected using the DAB staining kit (Komabiotech, Seoul, Korea). Additionally, to assess whether pLenti CMV SΔ19 serves as a more efficient expression vector compared to pcDNA3.1-SΔ19, the expression level of the spike protein was analyzed following the same aforementioned procedure.
Results
Construction of intron vector encoding SARS-CoV-2 spike protein
In order to investigate the potential of introns to enhance the expression of the SARS-CoV-2 spike protein, we initially cloned the C-terminal 19 amino acid-truncated D614G variant of the SARS-CoV-2 spike protein (derived from the Wuhan-Hu-1 strain) with a 1x HA tag into the EcoRI and NotI restriction sites of the pcDNA3.1 vector, resulting in the construction of pcDNA3.1-D614G S Δ19. Subsequently, an intron was inserted downstream of the CMV promoter, generating pcDNA3.1-intron-D614G S Δ19 (as depicted in Figure 1). Upon transfection of cells, two bands were observed in accordance with the anticipated sizes for the unprocessed spike protein (180 kDa) and the S2 subunit of the spike protein (90 kDa), as demonstrated in Figure 2. Furin cleavage of the spike protein into S1 (∼90–110 kDa) and S2 (∼80–100 kDa) results in a single observed band, due to their similar molecular weights, alongside a separate band representing the uncleaved protein. Notably, augmented expression of the SARS-CoV-2 D614G S Δ19 protein was detected in HEK293T cells transiently transfected with pcDNA3.1-intron-D614G S Δ19. To verify this finding, we compared the expression levels of SΔ19 in pcDNA3.1, both with and without an intron. As anticipated, the presence of introns resulted in increased expression levels of the SARS-CoV-2 S Δ19 protein.
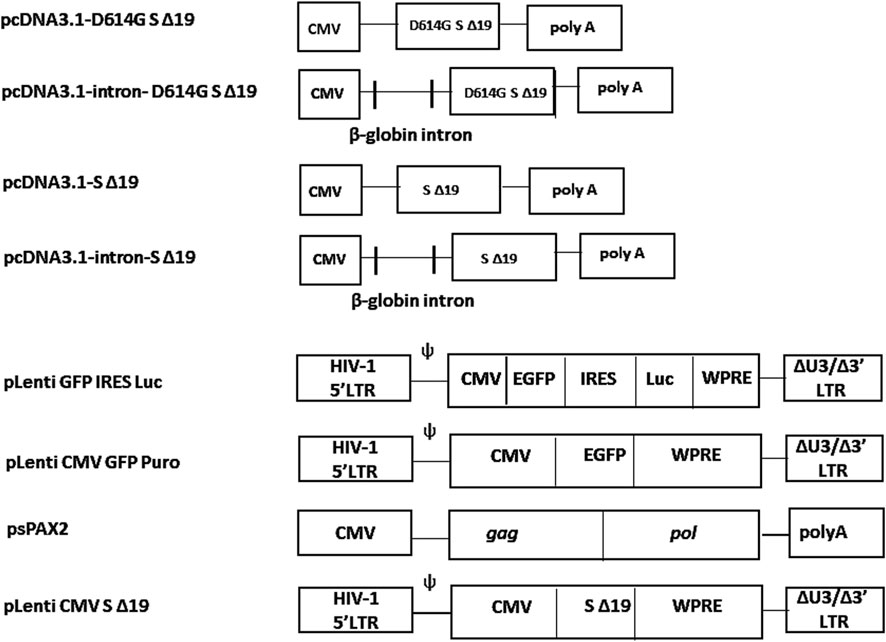
Figure 1. Structure of intron-containing vector and the WPRE-containg vectors. The plasmid pcDNA3.1-SΔ19 was modified to include the truncated cytoplasmic tail of the SARS-CoV-2 spike protein along with an HA tag. To generate pcDNA3.1-intron-SΔ19, the β-globin intron was inserted upstream of the SΔ19 sequence within pcDNA3.1-SΔ19. Plasmid pLenti CMV SΔ19 was created by introducing an XbaI-SalI fragment encoding the SARS-CoV-2 S Δ19 spike protein into the corresponding site of the pLenti CMV GFP Puro vector, which contains the WPRE element.
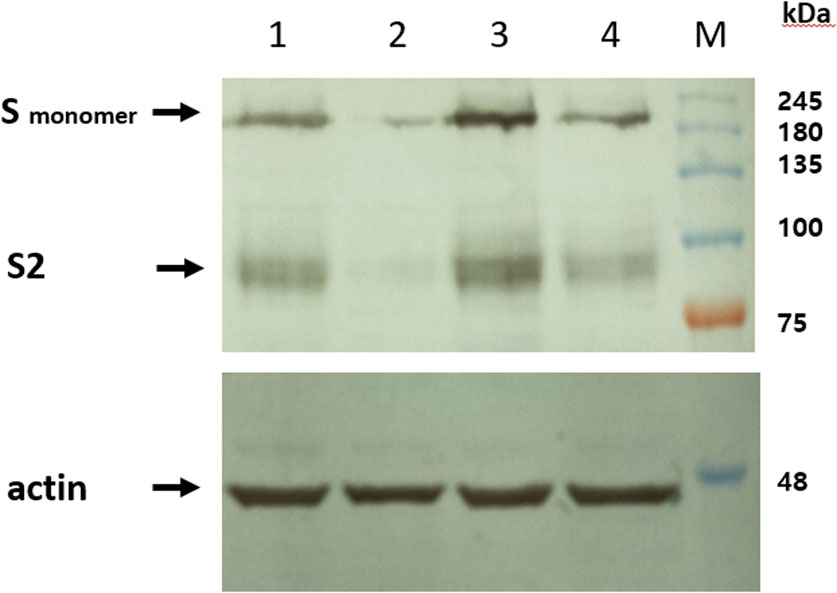
Figure 2. Intron vector increases expression of spike protein. HEK293T cells were transfected with pcDNA3.1 vectors containing intron sequences (pcDNA3.1-intron-D614G SΔ19 and pcDNA3.1-intron-SΔ19) as well as pcDNA3.1 vectors lacking introns (pcDNA3.1-D614G-SΔ19 and pcDNA3.1-SΔ19). The expression of the spike protein was assessed using western blot analysis. Lane 1, pcDNA3.1-intron-D614G SΔ19. Lane 2, pcDNA3.1-D614G-SΔ19. Lane 3, pcDNA3.1-intron-SΔ19. Lane 4, pcDNA3.1-SΔ19. A representative image from three independent experiments is presented.
Intron-containing vector enhances pseudotyping efficiency
Previous research has demonstrated that optimizing codons and utilizing vectors containing introns can enhance the expression of the spike protein of SARS-CoV (Moore et al., 2004; Schwegmann-Wessels et al., 2006). In our study, we employed the intron-containing pcDNA3.1 vector and achieved similar outcomes (Figure 2). In order to investigate whether increased expression could yield more of the pseudotyped lentiviruses, we developed a lentivirus system pseudotyped with the SARS-CoV-2 spike protein. As depicted in Figure 3, the titer of the viral vector produced from pcDNA3.1-intron-SΔ19 was three times higher than that generated from pcDNA3.1-SΔ19. The titers obtained from pcDNA3.1-intron-SΔ19, pcDNA3.1-SΔ19, and pCMV-VSV-G were 4.4 × 105 transducing units (TU)/mL, 1.3 × 105 transducing units (TU)/mL, and 1.2 × 106 transducing units (TU)/mL, respectively. To validate this finding, we conducted a luciferase assay. We constructed a lentiviral vector (pLenti GFP-IRES-Luc) capable of packaging and encoding both a GFP and luciferase reporter. The utilization of dual GFP and luciferase reporters facilitated the measurement of pseudovirus infectivity through microscopic observation (Figure 4A) or luciferase assay (Figure 4B). The luciferase assays revealed that the pseudovirus titer produced from pcDNA3.1-intron-D614G SΔ19 was two orders of magnitude higher than that generated from pcDNA3.1-D614G SΔ19. VSV-G and pseudoviruses lacking the spike protein were employed as positive and negative control viruses, respectively.
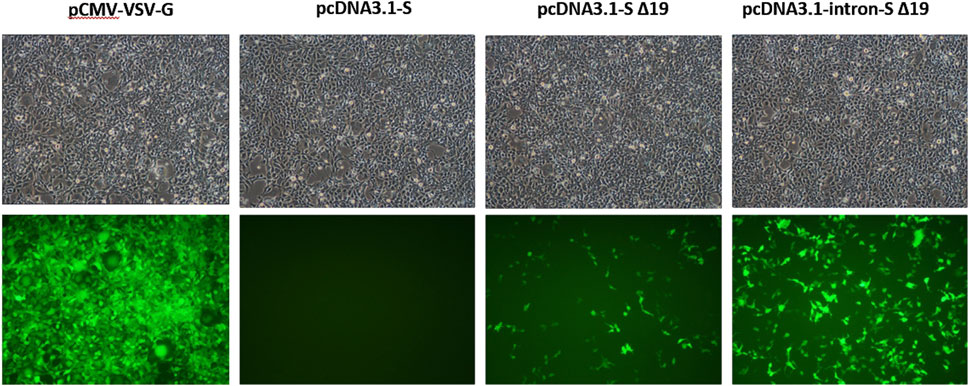
Figure 3. Comparison of the yields of pseudotyped virus generated from pcDNA3.1 with and without intron. HEK293T cells were cultured in 6-well plates and co-transfected with 1.2 µg of psPAX2, 1.6 µg of pLenti CMV GFP Puro, and 0.4 µg of one of the spike-expressing plasmids (pcDNA3.1-S, pcDNA3.1-SΔ19, and pcDNA3.1-intron SΔ19). The positive control, pCMV-VSV-G, was also included. HEK293-hACE cells in 12-well plates were then infected with 500 µL of pseudotype viruses obtained from the transfected HEK293T cells. The fluorescence images were captured from cells infected with pseudotyped viruses. The determination of pseudovirus titer is based on the quantification of GFP gene expression levels.
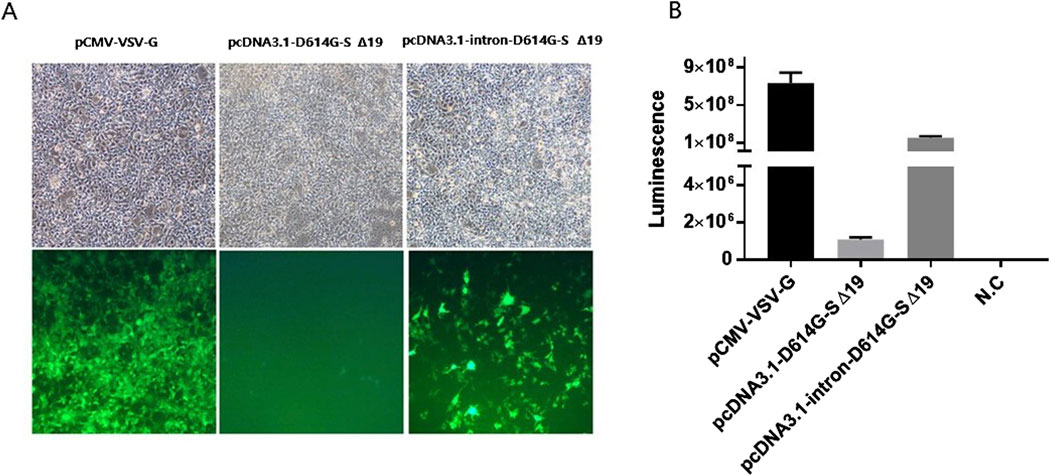
Figure 4. Titers of D614G spike-pseudotyped lentivirus in 293-hACE2 cells. (A) Microscopic images were obtained, showing the transduction of 293-hACE2 cells with D614G spike-pseudotyped lentivirus containing the GFP-IRES-Luciferase backbone. (B) The titers of the GFP-IRES-Luciferase backbone were determined by quantifying the relative luciferase units (RLUs) after infecting 2 × 104 293-hACE2 cells per well in 96-well plates with 100 µL of each pseudotyped virus. N.C; pseudotyped virus generated in the absence of spike is used as negative control. The experiments were performed in triplicates and repeated three times.
Construction of lentiviral vector encoding SARS-CoV-2 spike protein
Previous studies have demonstrated the utility of WPRE (Woodchuck Hepatitis Virus Posttranscriptional Regulatory Element) in enhancing gene expression after transcription (Higashimoto et al., 2007; Zufferey et al., 1999). In this study, we aimed to assess whether WPRE could enhance the expression of the spike gene delivered by the lentiviral vector, pLenti CMV S Δ19, which was derived from an HIV-based self-inactivating vector (pLenti CMV GFP Puro). Upon inserting WPRE downstream of the spike gene, pLenti CMV S Δ19 exhibited increased spike expression compared to pcDNA3.1-S Δ19 (Figure 5A). These findings indicate that WPRE has the potential to stimulate spike expression. To confirm the formation of spike-induced syncytia, pLenti CMV S Δ19 and pcDNA3.1-S Δ19 were transiently transfected into HEK293-hACE2 cells. Transient transfection with pLenti CMV S Δ19 resulted in even stronger syncytia formation and cell death at 48 h post transfection (Figure 5B). To assess the capacity of lentiviral transduction to induce syncytia formation, SARS-CoV-2 spike protein was transduced into HEK293-hACE2 cells using a lentiviral vector containing WPRE. Successful packaging of pLenti CMV S Δ19 was confirmed, resulting in syncytia formation (Figure 6B). Notably, in the absence of VSV-G, the truncated spike protein retained the ability to pseudotype HIV core particles and induce syncytia formation (Figure 6C). Conversely, the lentiviral vector pLenti CMV GFP Puro, employed as a negative control, did not induce syncytia formation (Figure 6A).
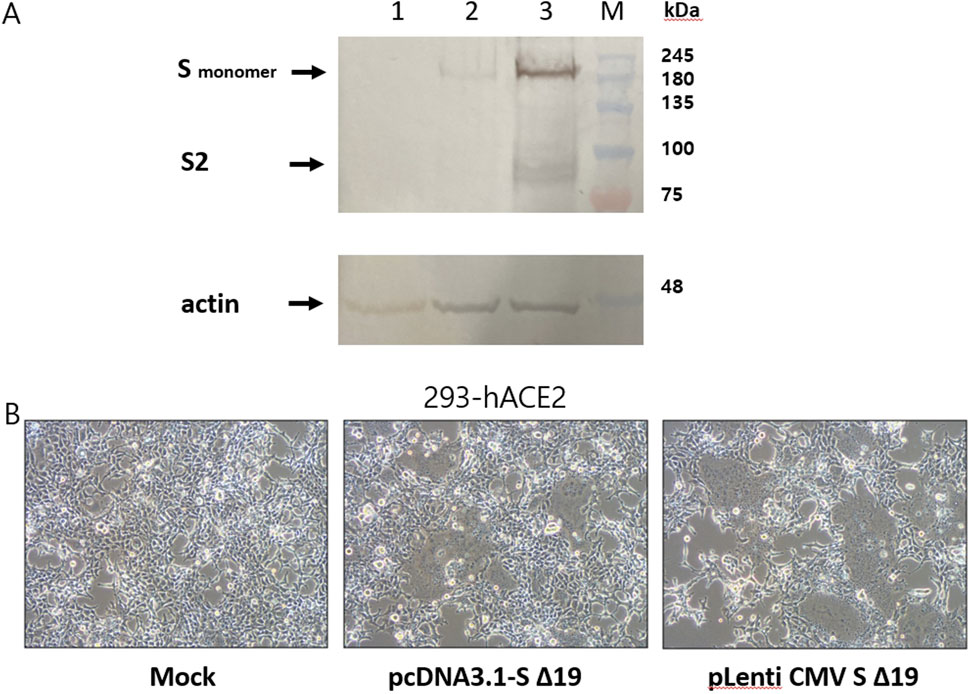
Figure 5. The WPRE-containing lentiviral vector increases expression of spike protein. (A) Lysates obtained from HEK293-hACE2 cells were subjected to Western blot analysis after transfection with either 1.0 µg of pcDNA3.1-SΔ19 or 1.0 µg of pLenti CMV SΔ19. Lane 1, HEK293-hACE2 cells transfected with pcDNA3.1. Lane 2, HEK293-hACE2 cells transfected with pcDNA3.1-SΔ19. Lane 3, HEK293-hACE2 cells transfected with pLenti CMV SΔ19. (B) To validate the expression of the spike protein, HEK293-hACE2 cells underwent transfection with either 1.0 µg of pcDNA3.1-SΔ19 or 1.0 µg of pLenti CMV SΔ19. The formation of syncytia was observed using a light microscope, and a representative image from three independent experiments is presented.
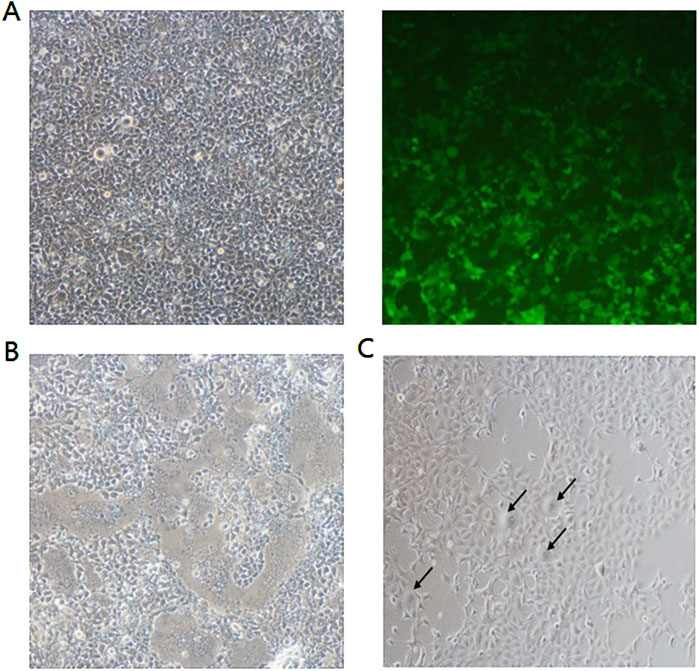
Figure 6. Syncytia formation was observed following transduction with lentiviral vectors expressing S Δ19. To determine the capacity of lentiviral transduction to induce syncytia formation, in addition to transient transfection, HEK293-hACE2 cells were infected with supernatants derived from HEK293T cells transfected with the following plasmids: (A) 1.2 µg psPAX2 + 0.4 µg pCMV-VSV-G + 1.6 µg pLenti CMV GFP Puro; (B) 1.2 µg psPAX2 + 0.4 µg pCMV-VSV-G + 0.4 µg pLenti CMV SΔ19; or (C) 1.2 µg psPAX2 + 0.4 µg pLenti CMV SΔ19. Syncytia formation was indicated by the black arrows and observed with light microscope. Magnification, ×100. A representative image of three independent experiments is shown.
Discussion
Previous studies have successfully developed pseudotyped viral systems for SARS-CoV and MERS-CoV (Giroglou et al., 2004; Grehan et al., 2015; Millet and Whittaker, 2016). Recently, researchers have created SARS-CoV-2 spike pseudotyped viruses utilizing HIV-1, murine leukemia virus (MLV), and VSV (Havranek et al., 2020; Lee et al., 2022; Yang et al., 2020). Among these, the HIV-1-based spike pseudotyped lentivirus offers a practical tool for screening SARS-CoV-2 entry inhibitors and conducting neutralization assays. To improve the titer of lentiviral vectors, it is recommended to truncate the spike protein cytoplasmic tail and introduce the D614G mutation. Although the D614G mutation enhances spike incorporation and pseudotyped virus titers, the effect is masked by cytoplasmic tail truncation (Yang et al., 2020; Yurkovetskiy et al., 2020; Chen et al., 2021). In this study, we developed an efficient expression vector to generate a high titer of SARS-CoV-2 spike-pseudotyped virus.
Previous studies have proposed that vectors containing β-globin introns could prevent mRNA degradation (Schwegmann-Wessels et al., 2006). Introns, which are noncoding sequences within genes, are excised during RNA splicing. Proper splicing facilitates the deposition of exon-junction complexes (EJCs), thereby enhancing mRNA stability and promoting efficient translation. As depicted in Figure 2, efficient expression of the spike protein can be achieved using vectors containing β-globin introns. The difference in band intensity observed between lane 2 and lane 4 in the SARS-CoV spike expression shown in Figure 2 is attributed to the distinct origins of the plasmid vectors. Specifically, pcDNA3.1-intron-D614G SΔ19 (lane 1) and pcDNA3.1-D614G SΔ19 (lane 2) were derived from pCAGGS-SARS-S-G614 (Addgene plasmid #156421), whereas pcDNA3.1-intron-SΔ19 (lane 3) and pcDNA3.1-SΔ19 (lane 4) were derived from GenScript (Piscataway, NJ, United States).To investigate whether an intron-containing vector can enhance pseudovirus titers, a SARS-CoV-2 spike-pseudotyped lentiviral vector system was developed. As shown in Figure 3, the inclusion of an intron-containing vector resulted in a more than three-fold increase in pseudotyped virus production. Additionally, we confirmed that removing the last 19 residues of the cytoplasmic tail improved pseudoviral titers. In addition to pseudoviruses packaging the GFP backbone, we also developed pseudoviruses packaging the GFP-IRES-Luciferase backbone. Consistent with the fluorescence microscope image of HEK293-hACE2 cells infected with pseudoviruses (Figure 4A), the pseudovirus with pcDNA3.1-intron-D614G SΔ19 exhibited a strong luminescence signal of 108 RLUs compared to the pseudovirus with pcDNA3.1-D614G SΔ19 (106 RLUs) (Figure 4B).
While introns are well-known for facilitating gene expression, the Woodchuck Hepatitis Virus Posttranscriptional Regulatory Element (WPRE) can also stimulate transgene expression. For instance, the insertion of WPRE into retroviral or lentiviral vectors has been shown to enhance transgene expression (Higashimoto et al., 2007; Zufferey et al., 1999). The WPRE is a cis-acting RNA element that interacts with host RNA-binding proteins to prevent transcript degradation. Furthermore, WPRE facilitates the recruitment of nuclear export factors, thereby enhancing mRNA transport from the nucleus to the cytoplasm. This increased export efficiency leads to elevated cytoplasmic mRNA levels, ultimately promoting higher protein synthesis. Our data demonstrated that WPRE stimulated the expression of spike genes lacking introns when delivered via HIV-derived vectors. The Western blot results shown in Figure 5A indicate that spike protein expression was elevated in cells transfected with the vector containing WPRE compared to those transfected with the vector lacking WPRE. However, this increase in spike protein expression due to WPRE did not significantly impact syncytial formation (Figure 5B), as no significant difference was observed in syncytial formation between pLenti CMV S Δ19 and pcDNA3.1-SΔ19. The syncytia formation demonstrated in Figures 6B, C further indicates that syncytia can arise not only through transient overexpression of the Spike protein (as shown in Figure 5B) but also through transduction with a lentiviral vector encoding the Spike protein. These results suggest that lentiviral vectors expressing S Δ19 may serve as a valuable tool for screening fusion inhibitors.
Although this study suggests that utilizing the β-globin intron and WPRE can increase the production yield of SARS-CoV-2 pseudovirus, these elements can also be applied to improving the yield of virus-like particles, optimizing viral antigen expression in viral vector-based vaccine development, and improving transgene expression in Adeno-associated virus (AAV) vectors to improve the efficacy of gene therapy.
In conclusion, our findings demonstrate that intron-containing vectors enhance spike protein expression and correlate with increased pseudotyped virus titers. Additionally, WPRE plays a role in boosting spike gene expression. Therefore, we recommend using a cytoplasmic tail-truncated spike protein in conjunction with an efficient expression vector to generate high-titer pseudotyped viruses.
Limitations of the study
To maintain experimental consistency, spike nucleotide sequences were identical across all plasmid constructs, excluding those encoding the D614G substitution and truncation variants; however, two differently codon-optimized spike plasmids were employed in this investigation. Additionally, a neutralization assay utilizing serum or antibodies should be conducted to confirm the full functionality of the pseudovirus generated in this study. Given that most studies employ pseudoviruses for neutralization assays, it is essential to validate the applicability of this novel method in the context of neutralization.
Data availability statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Author contributions
Y-TJ designed research and finalized the manuscript; HJ and YC performed research and analyzed data. All authors contributed to the article and approved the submitted version.
Funding
The author(s) declare that financial support was received for the research and/or publication of this article. This research was supported by “The Government-wide R&D to Advance Infectious Disease Prevention and Control,” Republic of Korea (grant number: HG23C161100). The Department of Microbiology was supported through the Research-Focused Department Promotion & Interdisciplinary Convergence Research Project as a part of the Support Program for University Development for Dankook University in 2024.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The authors declare that no Generative AI was used in the creation of this manuscript.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontierspartnerships.org/articles/10.3389/av.2025.14262/full#supplementary-material
References
Brinster, R. L., Allen, J. M., Behringer, R. R., Gelinas, R. E., and Palmiter, R. D. (1988). Introns increase transcriptional efficiency in transgenic mice. Proc. Natl. Acad. Sci. U. S. A. 85, 836–840. doi:10.1073/pnas.85.3.836
Buchrieser, J., Dufloo, J., Hubert, M., Monel, B., Planas, D., Rajah, M. M., et al. (2020). Syncytia formation by SARS-CoV-2-infected cells. EMBO J. 39, e106267. doi:10.15252/embj.2020106267
Cattin-Ortolá, J., Welch, L. G., Maslen, S. L., Papa, G., James, L. C., and Munro, S. (2021). Sequences in the cytoplasmic tail of SARS-CoV-2 Spike facilitate expression at the cell surface and syncytia formation. Nat. Commun. 12, 5333. doi:10.1038/s41467-021-25589-1
Chakraborty, C., Bhattacharya, M., and Sharma, A. R. (2021). Present variants of concern and variants of interest of severe acute respiratory syndrome coronavirus 2: their significant mutations in S-glycoprotein, infectivity, Re-infectivity, immune escape and vaccines activity. Rev. Med. Virol. 32, e2270. doi:10.1002/rmv.2270
Chavda, V. P., Patel, A. B., and Vaghasiya, D. D. (2022). SARS-CoV-2 variants and vulnerability at the global level. J. Med. Virol. 94, 2986–3005. doi:10.1002/jmv.27717
Chen, H. Y., Huang, C., Tian, L., Huang, X., Zhang, C., Llewellyn, G. N., et al. (2021). Cytoplasmic tail truncation of SARS-CoV-2 spike protein enhances titer of pseudotyped vectors but masks the effect of the D614G mutation. J. Virol. 95, e0096621. doi:10.1128/JVI.00966-21
Cheng, Y. W., Chao, T. L., Li, C. L., Wang, S. H., Kao, H. C., Tsai, Y. M., et al. (2021). D614G Substitution of SARS-CoV-2 spike protein increases syncytium formation and virus titer via enhanced furin-mediated spike cleavage. mBio 12, e0058721. doi:10.1128/mBio.00587-21
Conceicao, C., Thakur, N., Human, S., Kelly, J. T., Logan, L., Bialy, D., et al. (2020). The SARS-CoV-2 Spike protein has a broad tropism for mammalian ACE2 proteins. PLoS Biol. 18, e3001016. doi:10.1371/journal.pbio.3001016
Di Genova, C., Sampson, A., Scott, S., Cantoni, D., Mayora-Neto, M., Bentley, E., et al. (2021). Production, titration, neutralisation, storage and lyophilisation of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) lentiviral pseudotypes. Bio. Protoc. 11, e4236. doi:10.21769/BioProtoc.4236
Fu, X., Tao, L., and Zhang, X. (2021). Comprehensive and systemic optimization for improving the yield of SARS-CoV-2 spike pseudotyped virus. Mol. Ther. Methods Clin. Dev. 20, 350–356. doi:10.1016/j.omtm.2020.12.007
Giroglou, T., Cinatl, J., Rabenau, H., Drosten, C., Schwalbe, H., Doerr, H. W., et al. (2004). Retroviral vectors pseudotyped with severe acute respiratory syndrome coronavirus S protein. J. Virol. 78, 9007–9015. doi:10.1128/JVI.78.17.9007-9015.2004
González, J. M., Pénzes, Z., Almazán, F., Calvo, E., and Enjuanes, L. (2002). Stabilization of a full-length infectious cDNA clone of transmissible gastroenteritis coronavirus by insertion of an intron. J. Virol. 76, 4655–4661. doi:10.1128/jvi.76.9.4655-4661.2002
Grehan, K., Ferrara, F., and Temperton, N. (2015). An optimised method for the production of MERS-CoV spike expressing viral pseudotypes. MethodsX 13, 379–384. doi:10.1016/j.mex.2015.09.003
Havranek, K. E., Jimenez, A. R., Acciani, M. D., Lay Mendoza, M. F., Reyes Ballista, J. M., Diaz, D. A., et al. (2020). SARS-CoV-2 spike alterations enhance pseudo particle titers and replication-competent VSV-SARS-CoV-2 virus. Viruses 12, 1465. doi:10.3390/v12121465
Higashimoto, T., Urbinati, F., Perumbeti, A., Jiang, G., Zarzuela, A., Chang, L. J., et al. (2007). The woodchuck hepatitis virus post-transcriptional regulatory element reduces readthrough transcription from retroviral vectors. Gene Ther. 14, 1298–1304. doi:10.1038/sj.gt.3302979
Hoffmann, M., Kleine-Weber, H., Schroeder, S., Krüger, N., Herrler, T., Erichsen, S., et al. (2020). SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8. doi:10.1016/j.cell.2020.02.052
Hu, J., Gao, Q., He, C., Huang, A., Tang, N., and Wang, K. (2020). Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2. Genes Dis. 7, 551–557. doi:10.1016/j.gendis.2020.07.006
Johnson, M. C., Lyddon, T. D., Suarez, R., Salcedo, B., LePique, M., Graham, M., et al. (2020). Optimized pseudotyping conditions for the SARS-COV-2 spike glycoprotein. J. Virol. 94, e01062-20. doi:10.1128/JVI.01062-20
Lee, S. Y., Kim, D. W., and Jung, Y. T. (2022). Construction of SARS-CoV-2 spike-pseudotyped retroviral vector inducing syncytia formation. Virus Genes 58, 172–179. doi:10.1007/s11262-022-01890-z
Lei, C., Qian, K., Li, T., Zhang, S., Fu, W., Ding, M., et al. (2020). Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat. Commun. 11, 2070. doi:10.1038/s41467-020-16048-4
Lu, R. J., Zhao, X., Li, J., Niu, P. H., Yang, B., Wu, H. L., et al. (2020). Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395, 565–574. doi:10.1016/S0140-6736(20)30251-8
Millet, J. K., Tang, T., Nathan, L., Jaimes, J. A., Hsu, H. L., Daniel, S., et al. (2019). Production of pseudotyped particles to study highly pathogenic coronaviruses in a biosafety level 2 setting. J. Vis. Exp. 1, 10.3791–59010. doi:10.3791/59010
Millet, J. K., and Whittaker, G. R. (2016). Murine Leukemia Virus (MLV)-based coronavirus spike-pseudotyped particle production and infection. Bio. Protoc. 6, e2035. doi:10.21769/BioProtoc.2035
Moore, M. J., Dorfman, T., Li, W., Wong, S. K., Li, Y., Kuhn, J. H., et al. (2004). Retroviruses pseudotyped with the severe acute respiratory syndrome coronavirus spike protein efficiently infect cells expressing angiotensin-converting enzyme 2. J. Virol. 78, 10628–10635. doi:10.1128/JVI.78.19.10628-10635.2004
Moore, M. J., and Proudfoot, N. J. (2009). Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 136, 688–700. doi:10.1016/j.cell.2009.02.001
Nguyen, H. T., Zhang, S., Wang, Q., Anang, S., Wang, J., Ding, H., et al. (2020). Spike glycoprotein and host cell determinants of SARS-CoV-2 entry and cytopathic effects. J. Virol. 95, e02304-e02320. doi:10.1128/JVI.02304-20
Nie, J., Li, Q., Wu, J., Zhao, C., Hao, H., Liu, H., et al. (2020). Quantification of SARS-CoV-2 neutralizing antibody by a pseudotyped virus-based assay. Nat. Protoc. 15, 3699–3715. doi:10.1038/s41596-020-0394-5
Roy, S., Ghani, K., de Campos-Lima, P. O., and Caruso, M. (2021). A stable platform for the production of virus-like particles pseudotyped with the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) spike protein. Virus Res. 295, 198305. doi:10.1016/j.virusres.2021.198305
Schwegmann-Wessels, C., Ren, X., and Herrler, G. (2006). Intracellular transport of the S proteins of coronaviruses. Adv. Exp. Med. Biol. 581, 271–275. doi:10.1007/978-0-387-33012-9_45
Shang, J., Ye, G., Shi, K., Wan, Y., Luo, C., Aihara, H., et al. (2020). Structural basis of receptor recognition by SARS-CoV-2. Nature 581, 221–224. doi:10.1038/s41586-020-2179-y0.1038/s41586-020-2179-y
Thakur, N., Gallo, G., Elreafey, A. M. E., and Bailey, D. (2021). Production of recombinant replication-defective lentiviruses bearing the SARS-CoV or SARS-CoV-2 attachment spike glycoprotein and their application in receptor tropism and neutralisation assays. Bio. Protoc. 11, e4249. doi:10.21769/BioProtoc.4249
Ulper, L., Sarand, I., Rausalu, K., and Merits, A. (2008). Construction, properties, and potential application of infectious plasmids containing Semliki Forest virus full-length cDNA with an inserted intron. J. Virol. Methods 148, 265–270. doi:10.1016/j.jviromet.2007.10.007
Yang, R., Huang, B., Ruhan, A., Li, W., Wang, W., Deng, Y., et al. (2020). Development and effectiveness of pseudotyped SARS-CoV-2 system as determined by neutralizing efficiency and entry inhibition test in vitro. Biosaf. Health 2, 226–231. doi:10.1016/j.bsheal.2020.08.004
Yu, J., Li, Z., He, X., Gebre, M. S., Bondzie, E. A., Wan, H., et al. (2021). Deletion of the SARS-CoV-2 spike cytoplasmic tail increases infectivity in pseudovirus neutralization assays. J. Virol. 95, e00044-21. doi:10.1128/JVI.00044-21
Yurkovetskiy, L., Wang, X., Pascal, K. E., Tomkins-Tinch, C., Nyalile, T. P., Wang, Y., et al. (2020). Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183 (3), 739–751.e8. doi:10.1016/j.cell.2020.09.032
Zheng, Y., Larragoite, E. T., Williams, ESCP., Lama, J., Cisneros, I., Delgado, J. C., et al. (2021). Neutralization assay with SARS-CoV-1 and SARS-CoV-2 spike pseudotyped murine leukemia virions. Virol. J. 18, 1. doi:10.1186/s12985-020-01472-1
Keywords: β-globin intron, COVID-19, pseudovirus, Sars-CoV-2, WPRE
Citation: Jin H, Cho YR and Jung YT (2025) The effect of beta-globin intron and WPRE on transient expression of SARS-CoV-2 spike and generation of pseudotyped lentiviruses. Acta Virol. 69:14262. doi: 10.3389/av.2025.14262
Received: 26 December 2024; Accepted: 28 March 2025;
Published: 03 April 2025.
Edited by:
Boris Klempa, Slovak Academy of Sciences, SlovakiaReviewed by:
Jaroslav Holly, National Institute of Allergy and Infectious Diseases (NIH), United StatesBoris Klempa, Slovak Academy of Sciences, Slovakia
Copyright © 2025 Jin, Cho and Jung. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yong Tae Jung, eWp1bmdAZGFua29vay5hYy5rcg==
 Hyeop Jin
Hyeop Jin Yong Tae Jung
Yong Tae Jung